- class prepare.Preparation(input_file, logginglevel='info', external_logger=None, loggername=None)
-
Bases:
objectImport input parameters, and create input files.
- create_logger(logginglevel='info', loggername=None, external_logger=None)
-
Create and return a logger.
- generate_invers_input(local_date)
-
Fills the invers input file.
Calls get_inv_parameters with the local date.
- Parameters: local_date (dt.datetime) – the date in local time to be processed
- Returns: prf_input_files, skipped_spectra –
-
-
- prf_input_files (list): A list of paths to the input files.
-
- skipped_spectra (list): List containing all spectra skipped at this day, due to missing pressure values. This list is provided by get_spectra_pT_input called in get_inv_parameters.
-
generate_pcxs_input(local_date)-
Fills the pcxs input file.
Calls get_pcxs_parameters with local_date.
- Parameters: local_date (dt.datetime) – the date in local time to be processed
- Returns: prf_input_file (str) – Path to the input file.
-
- generate_preprocess_input(meas_date)
-
Fills the preprocess tempate file.
Calls self.get_prep_parameters(meas_date) :param meas_date: the date in measurement time to be processed :type meas_date: dt.datetime
- Returns: prf_input_file (str, or None) – If no igrams are found, return None, else the path to the input file
- get_coords()
-
Return dict of coords.
If coords were not given or contain None for at least one coordinate, the coord_file will be read. If the coord_file was also not given, operation will be terminated.
- get_coords_from_file(date)
-
Return the coordinates from the coord file.
- get_igrams(meas_date)
-
Search for interferograms on disk and return a list of files.
- get_ils(meas_date)
-
Return ILS parameters from measurement date.
The ILS parameters are either taken from the input file, from the ILS list. In case of processing a different instrument that the EM27/SUN, unity ILS parameters of ME=0.983 and PE = 0.0 are assumed.
- Parameters: meas_date (dt.datetime) – date in measurement time
- Returns: ME1, PE1, ME2, PE2 (tuple) – ILS parameters
get_ils_from_file(date)-
Read the ILS parameters form the given file.
- Parameters: date (dt.datetime) – If multiple ILS Parameters are given in the list, get the newest ILS parameters, that are already valid at date.
- Returns: ils_parameters (tuple) – MEChan1, PEChan1, MEChan2, PEChan2
get_inv_parameters(local_date)-
Return parameters to fill the invers20.inp template.
If spectra from two measurement days (i.e. different folders) belong to one local date, spectra_pT_input is a list with two elements, else, the it has one element.
For each element in the spectra_pT_input list a dict containing the parameters for the input files is generated.
The list of spectra belongig to one local date is sorted by the measurement date of the spectra which is determined by the function get_times_of(spectrum).
- Parameters: local_date (dt.datetime) – date in local time
- Returns: parameters, skipped_spectra –
-
-
- parameters (list): Contains one or two dict objects, depending if all spectra of the local date are stored in the same YYMMDD folder.
-
- skipped_spectra (list): List containing all spectra skipped at this day, due to missing pressure values. This list is provided by get_sepctra_pT_input.
-
- get_localdate_spectra()
-
Return dict linking all spectra to local dates.
- Returns: localdate_spectra (dict) – containing the a list of full pathes to the spectra for each local date in the format {local_date: [“path/YYMMDD_HHMMSSSN.BIN”, …]}
- get_mapfiles(local_noon_utc)
-
Return mapfiles of date and following date of the local noon in UTC.
- get_meas_dates(start_date=None, end_date=None)
-
Return a list of dates in measurement time for the site + instrument.
Truncate the list if start_date and end_date are given.
Parameters:
- start_date (dt.date) – optional start date
- end_date (dt.date) – optional end date
get_pcxs_parameters(local_date)-
Return parameters to fill the pcxs20.inp template.
- Parameters: local_date (dt.datetime) – date in local time
- Returns: parameters (dict) – dict containing the parameters to fill the pcxs template.
- get_prep_parameters(meas_date)
-
Return Parameters to be replaced in the preprocess template.
- Parameters: meas_date (dt.datetime) – date in measurement time
- Returns: parameters (dict) – dict with parameters to fill the preporcess template.
- get_prf_input_path(template_type, date=None)
-
Return path to the corresponding prf_input_file.
- get_spectra(meas_date)
-
Return list of spectra for a given date (in measurement time).
- Parameters: date (dt.datetime) – in measurement time
- Returns: spectra (list) – with full path to all spectra of measurement date [“path_to/YYMMDD_HHMMSSSN.BIN”, …]
get_spectra_pT_input(local_date)-
Return invers formatted pT infos for given local date.
If two measurement dates belong to one local date, spectra_pT_input contains two lists. The list is split based on the filename of the spectra, since the filename refers to measurement time.
The string has the format YYMMDD_HHMMSSSN.BIN, pressure, T_PBL
This function replaces the pt_intraday.inp file from older PROFFAST versions. Note that T_PBL is currently set to 0.0.
- Parameters: local_date (dt.datetime) – Date in local time
- Returns: spectra_pT_input (list), skipped_spectra (list) –
-
-
- spectra_pT_input: List containing a list of strings with spectra and pT infos.
-
- skipped_spectra (list): List containing all spectra skipped at this day, due to missing pressure values.
-
- get_template_path(template_type)
-
Return path to the corresponding template file.
- interpolate_map_files(local_date)
-
Interpolate GGG2020 map files.
Generate a map file at 12:00 local time. This method is only called for mapfiles of type GGG2020. The mapfile is created in <result_folder>/interpolated_mapfiles with the following filename:
-
“<site_abbrev><local_noon_utc>_Z.map”
The folder interpolated_mapfiles is created in this function.
- Parameters: local_date (dt.datetime) – datetime in local time
-
prepare_map_file(local_date)-
Generate map file if GGG2020 map file are used.
- Parameters: local_date (dt.datetime) – date in local time
- Returns: success (bool) – True if map files were found and created False if no files were found.
-
- replace_params_in_template(parameters, template_type, prf_input_file)
-
Generate a site specific input file by using a template.
- Parameters:
-
-
parameters (dict) – Containing keys which match the variable names in the template file. They are replaced by the entries.
-
template_type (str) – Can be “prep”, “pt”, “inv” or “pcxc”
-
prf_input_file (str) – The filename of the input file
-
class prepare.PylotOnly(name='')-
Bases:
FilterA filter which filters out all logs not originating from the pylot.
This is used when an external logger is provided, to prevent external logging messages to show up in the PROFFASTpylot custom log file.
- filter(record)
-
Determine if the specified record is to be logged.
Returns True if the record should be logged, or False otherwise. If deemed appropriate, the record may be modified in-place.
PROFFASTpylot v1.3 documentation
Measurements of atmospheric greenhouse gas (GHG) concentrations are important to assess the effect of climate change mitigation policies. Additionally, climate models depend on a precise knowledge of greenhouse gas abundances and emissions. A variety of measurement methods is addressing these needs. The Collaborative Carbon Column Observing Network (COCCON) was established in 2019, as a supporting framework for users of the portable Fourier-Transform spectrometers EM27/SUN, that measures precisely and accurately GHG column abundances from near-infrared solar absorption spectra. To ensure common quality standards across the COCCON, raw EM27/SUN measurements are processed with the PROFFAST Fortran routines. The Python interface PROFFASTpylot significantly reduces the workload during the processing of large sets of observational data and supports a network-wide consistent data processing.
For more information about PROFFAST, see Data Processing. The PROFFASTpylot source code is available at EUDAT GitLab.
If you have any comments or questions, contact Darko Dubravica or Frank Hase. You are welcome to contribute.
This documentation and the PROFFASTpylot source code is licensed under GPL-3.0.
©2023, Lena Feld, Benedikt Herkommer, Karlsruhe Institut of Technology.
User Guide
Getting started: Installation & Updates, Usage, Pressure Input, Folder Structure
User Information: List of all Input Parameters, Time Offsets, ILS Parameters, Instrument Parameters, Advanced Logging Options, Troubleshooting
Developer Information: Contribution Notes, Developer Guide, Modules (Pylot, Filemover, Prepare, Pressure)
Getting Updates
If you used git during installation, you can easily get updates by entering
git pull
in a git bash or in a Terminal in your proffastpylot folder. This command will download all available updates. If you downloaded PROFFASTpylot as zip file, you need to redo all steps of this installation script.
PROFFASTpylot - Installation
Installation - Content
-
Prerequisites
-
Download the PROFFASTpylot repository
-
Get PROFFAST and copy it to
proffastpylot -
Create a virtual environment in python
-
Install PROFFASTpylot
-
Resulting folder structure
-
Test the installation by running an example dataset
-
Getting Updates
1. Prerequisites
For using PROFFASTpylot you need Python 3.7 or newer. The PROFFAST and PROFFASTpylot can be used in Windows and Linux environments. A step-by-step installation instruction for both environments is given in the following. We did not test the software for Mac environments.
2. Download the PROFFASTpylot repository
Clone the PROFFASTpylot repository using git
We recommend downloading the files using git (https://www.git-scm.com).
It will make future updates easier.
git clone https://gitlab.eudat.eu/coccon-kit/proffastpylot.git
A folder proffastpylot containing all program files will be created.
Alternatively: Download the PROFFASTpylot repository as a zip file
If you don't want to use git, you can instead download and unpack the zip file https://gitlab.eudat.eu/coccon-kit/proffastpylot/-/archive/master/proffastpylot-master.zip
Extracting it will create the folder proffastpylot.
3. Get PROFFAST and copy it to the proffastpylot folder
Download PROFFAST
Download PROFFAST Version 2.4 from the KIT website:
https://www.coccon.kit.edu/69.php
Compile PROFFAST (only Linux)
For Windows users, the executables are already provided, on Linux systems you need to create them from source.
If not present on your system, first install the gfortan compiler.
Secondly, run the installation script for compilation from the prf folder.
cd prf/
bash install_proffast_linux.sh
Copy the prf directory
Copy the prf folder that was extracted from the zip file into proffastpylot.
4. Create a virtual environment in python
We recommend using a virtual environment (venv) to avoid conflicts between any other packages or Python modules.
-
(Only first time) Navigate to the
proffastpylotfolder using a terminal. -
(Only first time) Enter
python -m venv prf_venv.
This command will create a folder namedprf_venvwhich contains the virtual environment -
Activate the virtual environment every time you run PROFFASTpylot with
-
Windows PowerShell:
.\prf_venv\Scripts\Activate.ps1 -
Windows Commandline:
.\prf_venv\Scripts\activate -
Linux:
source prf_venv/bin/activate
-
-
To deactivate the virtual environment you can run
deactivate
Note that all packages to be installed with pip install will only affect the virtual environment and not the local Python installation.
In case of a problem, take a look at the Troubleshooting article of this documentation.
You need to activate the virtual environment before each run of PROFFASTpylot by executing the command in step 3, the other steps need to be executed only the first time.
5. Install the PROFFASTpylot repository
Activate the virtual environment (see above).
Navigate to proffastpylot and enter
pip install --editable .
6. Resulting folder structure
If you follow exactly the installation guide your folder structure should look like the following:
proffastpylot
├── prf_venv
│ ├── ...
├── docs
│ ├── ...
├── example
│ ├── input_sodankyla_example.yml
│ ├── log_type_pressure.yml
│ └── run.py
├── prf
│ ├── docs
│ ├── inp_fast
│ ├── inp_fwd
│ ├── preprocess
│ ├── source
│ ├── out_fast
│ └── wrk_fast
├── prfpylot
│ ├── ...
└── setup.py
7. Test the installation by running an example dataset
To test the installation, we provide example raw data and a reference result file to compare the file to. The example can be executed by navigating to the example folder and execute python run.py (please ensure that your virtual environment is activated).
When first running the program, it will ask you to download the example file data to your local computer.
After the run is complete, please compare your results to the data given in example\Reference_Output_Example_Sodankyla.csv. The deviations should be less than 0.1 ppm for XCO2, 0.1 ppb for XCH4 and 0.1 ppb for XCO.
8. Getting Updates
If you used git during installation, you can easily get updates by entering
git pull
in a git bash or in a Terminal in your proffastpylot folder. This command will download all available updates. If you downloaded PROFFASTpylot as zip file, you need to redo all steps of this installation script.
PROFFASTpylot - Usage
Usage - Content
-
General use
-
Input file creation
-
Starting the run
-
-
Special case: Process spectra directly
-
Folder Structure
1. General use
Ready-to-use Example
You can follow the usage of PROFFASTpylot with the help of an example from Sodankylä which is provided as example/run.py. The example input data (i.e. the example interferogram, map- and pressure files) are downloaded automatically when running run.py the first time. The runscript needs to be executed inside the example folder.
Executing PROFFAST with the PROFFASTpylot takes two steps
-
Create an input file with the required information
-
Execute PROFFASTpylot via a Python script
Both steps will be explained in more detail, in the following.
Input file creation
The input file (stored in the yaml format) contains all the key information required by PROFFASTpylot and PROFFAST, e.g., the location of the input and output files, or meta-data about the data to be processed. An example with explanations (example_sodankyla_input.yml) is provided. It contains all options that are required to process the example data set. Adjust this file to your requirements.
Starting the processing
For starting the processing, you need to create an instance of the Pylot class with an input file.
from prfpylot.pylot import Pylot if __name__ == "__main__": input_file = "input_sodankyla_example.yml" MyPylot = Pylot(input_file=input_file)
Note that the if __name__ == "__main__" statement needs to be placed before initializing Pylot to prevent problems with the multiprocessing on Windows.
Afterwards all steps of PROFFAST can be executed automatically one after the other:
MyPylot.run(n_processes)
Alternatively, you can run all steps of PROFFAST individually with the following commands:
n_processes = 2 try: MyPylot.run_preprocess(n_processes) MyPylot.run_pcxs(n_processes) MyPylot.run_inv(n_processes) MyPylot.combine_results() finally: MyPylot.clean_files()
You can execute run.py to test this with the example data provided.
Alternative way of starting the processing
In the case above, we loaded a yaml file to get the input parameters. However, when processing larger datasets or embedding PROFFASTpylot into an larger environment, it can be usefull to hand over the input parameters directly as a dictionary instead of a file.
In this case the initialization would look like:
from prfpylot.pylot import Pylot input_dict = { "instrument_number": "SN039", "site_name": "Sodankyla", "site_abbrev": "so", # ... and all other needed parameters } if __name__ == "__main__": MyPylot = Pylot(input_file=input_dict)
2. Special case: Process spectra directly
If the spectra are already available, set the option start_with_spectra to True in the input file. The path to the spectra is given to PROFFASTpylot by the entry analysis_path. Note, that the folder analysis_path must have the following substructure: analysis/SiteName_InstrumentNumber/YYMMDD.
Afterwards, Pylot.run() will not execute preprocess.
3. Folder structure
The results results will be created automatically. Please see the Folder structure article about how the results are organized.
PROFFASTpylot - Pressure Input
The pressure input was reorganized in version 1.1
This article explains how to handle pressure data with PROFFASTpylot. To perform the retrieval, PROFFAST needs pressure data from the measurement site. In PROFFAST 2.2 or newer, the pressure is read together with the spectra in the input file of invers. A template for this file can be found in prfpylot/templates. The pT_intraday.inp file is deprecated, the interpolation of the pressure is handled by PROFFASTpylot.
Provided options in PROFFASTpylot
Two parameters in the main input file specify how the pressure is handled by PROFFASTpylot.
-
pressure_pathis the location of the pressure files -
pressure_type_filelinks to a second input file in which the format of the pressure files is defined
Pressure type file
An example pressure type file is provided in example/log_type_pressure.yml, describing the KIT-style data format. The path to the pressure files recorded by the KIT datalogger need to be given as pressure_path.
To adapt this file two your own file format, the options are explained in the following.
Filename parameters
The filename parameters define how the filename of the pressure file is constructed. The pressure module of PROFFASTpylot will search for files with the naming <basename><time><ending>.
filename_parameters:
basename: ""
time_format: "%Y-%m-%d"
ending: "*.dat"
Dataframe parameters
In the dataframe parameters the internal formatting of your files is specified.
dataframe_parameters:
pressure_key: "BaroTHB40"
time_key: "UTCtime___"
time_fmt: "%H:%M:%S"
date_key: "UTCdate_____"
date_fmt: "%d.%m.%Y"
datetime_key: ""
datetime_fmt: ""
csv_kwargs:
sep: "\t"
The pressure file will be read in the following way.
import pandas as pd
params = filename_parameters
csv_kwargs = dataframe_parameters["csv_kwargs"]
filename = "".join(
[
params["basename"],
date.strftime(params["time_format"]),
params["ending"]
]
)
df = pd.read_csv(filename, **csv_kwargs)
For the date- and timestamp the datetime can be constructed from two separate columns (time_key and date_key) or one column (datetime_key). It will be parsed with the corresponding format string. In addition to the formats supported by the datetime package (see below) the key POSIX-timestamp can be used. This assumes the datetime column to be in seconds passed since the 1979-01-01 in UTC. df[pressure_key] should contain the corresponding pressure values.
For more information you can look at the pandas documentation of read_csv() and the datetime package.
Additional Options
-
A UTC offset of the pressure file can be given as
utc_offset. -
In
data_parametersyou can define minimum and maximum pressure values. -
The pressure values are multiplied by the
pressure_factor. It can be used to correct for a height offset or a different unit. The pressure is expected to be given in hPa. -
The
pressure_offsetis added to the pressure values. The pressure value is expected to be given in hPa. -
The
max_interpolation_timeis the maximal timely distance between two pressure data points. If the distance between two points is larger, the spectra belonging to the corresponding time is skipped. Can be set to a value in hours. Its default value is 2 hours. If the time is outside the range of the given pressure values, the nearest value will be used up to a time difference ofmax_interpolation_time.
PROFFASTpylot - Folder Structure
With PROFFASTpylot most of the folder structure can be chosen freely. This article gives an overview about the needed folders and files. They can be divided into four categories:
-
Input data: Interferograms, pressure files, map files
-
Output data: Spectra, results of the processing, config files
-
Program Files: This includes the folder to the PROFFAST binaries as well as the
proffastpylotdirectory (see note below). -
Steering Files: input files and run files for specific runs
All paths can be chosen freely (e.g. input data can be located on an external hard disk).
-
We recommend disentangling input data, output data and the program execution files from each other.
Note: To run the example files using run.py PROFFAST must be located inside the proffastpylot directory as described in the Installation article.
1. Input data
You can specify three paths for your input data:
-
Path of the interferograms
-
Path of the the pressure files
-
Path of the map files (containing atmospheric information).
Optionally a file with the coordinates of the measurements can be provided.
In our example all these folders and files are in example/input_data.
Interferograms
The interferogram_path should be a folder with the interferograms inside. It needs to have the following structure:
interferogram_path
├── YYMMDD
│ ├── YYMMDDSN.XXX
│ ├── YYMMDDSN.XXX
│ └── ...
├── YYMMDD
│ ├── ...
├── ...
Pressure input
Inside the pressure_path pressure measurements for each measurement day should be provided. The frequency of the files (e.g. monthly or daily) and the formatting is flexible. In the pressure_type_file the file format can be defined. See the Pressure Input article for how the pressure type file is organized.
Map Files
PROFFASTpylot supports the usage of the GGG2014 and GGG2020 map files. It detects automatically what kind of map file is used. The path to the files is specified by the map_path in the input file.
2. Output data
Two output paths have to be specified: the analysis_path and the result_path.
Analysis path
Day-specific output files are written to the analysis folder by PROFFAST automatically. The analysis folder is created with the following structure:
analysis_path
├── <site_name>_<instrument>
| ├──YYMMDD
│ | └── cal
| ├──YYMMDD
...
Inside the YYMMDD folder the spectra, which are named YYMMDD_HHMMSSSN.BIN (or SM.BIN) will be located. Note that this timestamp, as well as the folder name, correspond to the measurement time. The UTC time of the measurement, that is derived from the variable utc_offset in preprocess, is written to the header of the spectrum. For more information on time offsets, see the Time Offsets article.
The files that were located in YYMMDD/pT are handled elsewhere since version 1.1.
Result path
In the result folder run-specific information is stored. In the folder the retrieval results of all days are stored. Those files are merged to a single file comb_invparms_<site_name>_<instrument_nr>_<YYMMDD>-<YYMMDD>.csv. Furthermore, the logfiles of the runs are stored in result_path/logfiles.
3. Program Files
In principle the proffastpylot directory can be located elsewhere than the PROFFAST binaries. The location can be given as proffast_path in the input file. The default is proffastpylot/prf
4. Steering Files
The input files and Python scripts that call proffastpylot do not need to be located inside the proffastpylot directory.
Example for a possible folder structure
An example how a folder structure could look like is given below (most sub-directories are created automatically).
E:\EM27_Sodankyla_RawData
├── interferograms
├── map
└── pressure
D:\EM27_OutputData
├── analysis
│ └── Sodankyla_SN039
│ └── cal
└── results
└── Sodankyla_SN039_170608-170609
├── logfiles
├── ...
└── comb_invparms_Sodankyla_SN039_170608-170608.csv
D:\Sodankyla_Retrieval
├── run_sodankyla.py
├── sodankyla_pressure_type.yml
├── sodankyla_coords.csv
└── input_sodankyla.yml
D:\proffastpylot (does not contain any personal data)
├── docs
├── prf
│ ├── pxcs20.exe
│ ├── invers20.exe
│ ├── ...
│ ├── preprocess
│ │ └── preprocess4.exe
│ ├── out_fast
│ ├── inp_fast
│ ├── ...
│ └── wrk_fast
└── prfpylot
└── ...
In this example the relevant paths given in the input file would be:
coord_file: sodankyla_coords.csv
interferogram_path: E:\EM27_Sodankyla_RawData\interferograms
map_path: E:\EM27_Sodankyla_RawData\map
pressure_path: E:\EM27_Sodankyla_RawData\pressure
pressure_type_file: sodankya_pressure_type.yml
analysis_path: D:\EM27_OutputData\analysis
result_path: D:\EM27_OutputData\results
Note that paths can also be relative.
PROFFASTpylot - List of all Input Parameters
The example input file input_sodankyla_example.yml contains the most relevant input parameters to run the PROFFASTpylot. However, there are several more parameters which can be used optionally. In this file all parameters are given and explained.
Site information
-
instrument_number
The serial number of your EM27/SUN. -
site_name
The name of the current measurement site. -
site_abbrev
A two-letter abbreviation to assign your site to a map file. -
coords
Coordinates of the measurement. Alternatively, a coordinate file can be used (see below).-
lat: Latitude, degrees north in decimal format
-
lon: Longitude, degrees east in decimal format
-
alt: Altitude, km over sea level
-
-
coord_file
Give the coordinates in a comma separated file. An example file can be found inexamples/input_data/coords.csv -
utc_offset
The UTC-offset of your measurements. This can either be due to the measurement in a different time zone or due to an error in the syncing of the PC clock. -
note
Optional comment included in bin-files by PROFFAST preprocess.
Steering of the behavior:
-
min_interferogram_size
File size in MegaBytes. All interferograms smaller than the given size are skipped. -
igram_pattern
The file name pattern of the interferograms can be used to process only specific files in the interferogram folder. The default value is*.*, which means that all files in the interferogram folders are used for processing. -
start_with_spectra
TrueorFalse
Start the processing chain with already available spectra. The spectra have to be located at<analysis>/SiteName_InstrumentNumber/YYMMDD/calwhere<analysis>is the given analysis path. -
start_date
Date with the formatYYYY-MM-DD.
The first date to be processed. If not given, the earliest available date is the start date. -
end_date
Date with the formatYYYY-MM-DD.
The last date to be processed. If not given, the latest available date is the end date. -
delete_abscosbin_files
default:False
The*abscos.binfile contains the simulation of the atmosphere, which is the result of the PROFFAST pcxs program. Pcxs can then be skipped if the abscos files are present from a previous run. IfTruethe*abscos.binwill be removed after the run; otherwise they are kept inprf/wrk_fast. If the abscos files are not deleted, the pT and VMR files are copied instead of moved to the result folder in order to be present if PCXS is skipped in a following run. -
delete_input_files
default:False
IfFalse: The output of the PROFFAST programs will be moved to results folder. -
instrument_parameters
default:em27
To evaluate EM27/SUN data this parameter does not have to be given explicitly.
Possible values are:em27, tccon_ka_hr,tccon_ka_lr,tccon_default_hr,tccon_default_lr,invenio,vertex,ircubeor a path to an instrument-config file.
For more details refer to the Instrument Parameters article of the documentation. -
mapfile_wetair_vmr
default:None(determined during runtime)
If you are using other mapfiles than the standard ggg2020 or ggg2014 files you can set if the columns are based on wet air (True) or dry air (False). -
custom_mapfile
default:None.
This parameter can be used in two ways:-
Give a format specifier of the map-files you want to use. The names of the map-files must include a date string inclduing the year, month, day and the hour of the file.
If your mapfiles are, for example, named like “MyCustomMapFiles_YYYYMMDDHH_lat_long.map” give “MyCustomMapFiles_%Y%m%d%H*.map”. -
Give the path to a single file. Then, for all days of the processing this individual map-file is used. Note, that this not usefull for operational processing, but can be usefull for testing purposes.
-
-
use_measured_pressure_for_pcxs
default:False.
Normally, PROFFASTpcxs uses the surface pressure retrieved from the model data. When setting this parameter toTrue, PROFFASTpcxs uses the pressure measured at local noon instead.
Path settings
-
proffast_path
The path pointing to the PROFFAST folder. -
interferogram_path
The path to the interferograms. The data structure must be likeYYMMDD/YYMMDD*.XXX -
map_path
The path pointing to the map files. -
pressure_path
The path pointing to the pressure data. -
pressure_type_file
The path pointing to the pressure setup file. For more explanation see the article about pressure input in the documentation. -
analysis_path
In the analysis path the spectra produced by preprocess are written to. We recommend calling it an analysis. Within this folder, the PROFFASTpylot will create a folder structure likeanalysis/SiteName_InstrumentNumber/YYMMDD.Consult also the article about folder structure in the documentation. -
result_path
The merged results, the log files and optionally the input files are written to the result path. Within it, PROFFASTpylot will create a folder namedSiteName_InstrumentNumber_StartDate-EndDate.
PROFFASTpylot - Time Offsets
PROFFASTpylot allows to process measurements which have been recorded near the dateline, where one local day belongs to two different UTC days and vice versa.
Three different times are of importance
-
The UTC time
-
The measurement time (i.e. the time of the laptop)
-
The local time
with the corresponding offsets between them
utc to local time = utc to measurement time + measurement time to local time
The measurement time can be UTC time, local time, or any other time. The user has to provide the offset between UTC and measurement time in the input file (utc_offset). PROFFASTpylot calculates the local time automatically from the given coordinates. In case your computer clock was not accurate, utc_offset can also be set to non-integer values.
In case of UTC measurements near the date line it splits the spectra of one local day into two independent processes for the corresponding location of the spectra in the folders that are sorted by measurement date. This is necessary since the map file corresponds to the local noon.
Nevertheless, we strongly recommend you to use local time in case you are measuring near the date line. It makes handling of the data less confusing. E.g. the start- and end-date in the input file correspond to measurement time, not to local time.
PROFFASTpylot - ILS Parameters
The ILS parameters are distributed together with PROFFASTpylot in the file prfpylot/ILSList.csv. The correct parameters corresponding to the measurement time and instrument are read in automatically during runtime. We recommend to use these official ILS parameters. In special cases you can use individual ILS parameters in the input file.
The ILSList.csv file will be updated in regular intervals, make sure to keep your version of PROFFASTpylot up to date.
PROFFASTpylot - Instrument Parameters
WARNING: The settings described here are only for advanced users. For a regular processing of EM27/SUN spectra, the settings given here are not needed.
Starting with PROFFAST 2.3. there is the possibility to process data from other instruments than the EM27/SUN.
This is solved by adding new parameter in the PREPROCESS input file, which allows to adapt the behavior of the PREPROCESS to the instrument.
- PROFFASTpylot comes with several templates prepared already for the following instruments:
-
-
em27 -
tccon_ka_hr -
tccon_ka_lr -
tccon_default_hr -
tccon_default_lr -
invenio -
vertex -
ircube
-
To use one of these instruments set the parameter instrument_parameters to one of the options above.
Alternatively, it is possible to provide the path to a so called instrument-parameter-file. This option is only recommended to be used by expert-users. Therefore, no template is provided for this case. If you want to create your own instrument-parameter-file, please adapt one of the prepared parameter files. They are located in prfpylot\templates\instrument_templates.
PROFFASTpylot - Advanced Logging Options
The PROFFASTpylot has a built-in logging functionality. By default, the user do not have to care about this. This section is only intended for advanced usage of the PROFFASTpylot, e.g., when it is embedded into a larger environment.
The PROFFASTpylot uses the standard Python logging module. There are three possible use cases on how to use the logging functionality of PROFFASTpylot:
-
- Default mode:
-
As the PROFFASTpylot is mainly intended to be used as a standalone program, logging is configured such, that a custom logger object is created and stream and file handlers are added to this logger. The logger instance is initialized using the current datetime in the format YYMMDDHHMMSSssss as the logger’s name. In this mode the logger is mainly encapsulated to the PROFFASTpylot and difficult to be accessed from the outside.
-
- Submodule mode:
-
The
submodule modeis designed to use the Pylot as a submodule. This means that the logger instance is created outside of the PROFFASTpylot and passed to the Pylot as an argument. For this, thePylotclass has the argumentexternal_loggerto which an instance of the external logger is passed to. This mode has the advantage, that it is possible to use the logging before an instance of the Pylot has been created.
Note, that to this instance a logging file handler will be added, which results in the standard logging file within the result directory. To this handler, a filter is applied that only passes messages originating from the Pylot. Hence, the logfile in the PROFFASTpylot result directory does NOT contain any of the log messages from outside of the PROFFASTpylot. Furthermore, no stream handler is added. This must be taken care of outside of the PROFFASTpylot.
-
- Mainmodule mode:
-
The
mainmodule modeis designed to use the Pylot as the main module which takes care of the logging and also is used to record the logging messages of other modules. This is realized by passing a name to the Pylot using theloggernameargument, which will be used to initialize the logger. Then, the logger can be accessed by the code snippet below. This only works if the name of the logger passed to the Pylot is the same as the one used to initialize a new logger.
import logging
from prfpylot.pylot import Pylot
MyPylot = Pylot(input_file, logginglevel="info", loggername="my_logger")
my_logger = logging.get_logger("my_logger")
my_logger.info("This message is written from outside of the PROFFASTpylot")
All messages you will log there will also appear in the log file of the PROFFASTpylot.
PROFFASTpylot - Troubleshooting
In case of problems, please first take a look in the results/<site>_<instrument>_<dates>/logfiles folder. There, the output of PROFFASTpylot, as well as the output of the subroutines preprocess, pcxs and invers are stored. Sometimes the subroutines fail without returning an error; therefore, it looks successful at first in the pylot logfile.
If you encounter any problems, please do not hesitate to contact us via email or directly in GitLab. We will update this documentation with the occurring problems.
Installation
Problems creating a virtual environment
If you have several python installations, and some are not included to your PATH variable, it might happen that the creation of a virtual environment results in Error: [WinError 2] The system cannot find the file specified.
In this case the solution can be to give to full path to your Python executable whilst creating the virtual environment: C:\path\to\your\python\python.exe -m venv prf_venv
Execution
Infinite Loop: General logfile is being used by another process
Ensure if your run-script contains if __name__ == "__main__". If this is missing, it can crash the multiprocessing in a Windows environment.
PROFFASTpylot - Contribution Notes
If you have any idea for improvement, find a bug, or if you are missing a feature, you are welcome to contribute. PROFFASTpylot already improved a lot because of feedback we received.
Please check first if your idea was implemented already in the dev branch. If not, you can either
-
contact us directly via email,
-
create an issue at Gitlab or
-
implement the solution and make a merge request pointing to the
devbranch.
We are happy about every contribution.
Developer Guide
This document gives an overview about the structure of the PROFFASTpylot code. It should help all users who want to adapt PROFFASTpylot to their own needs.
The general structure of the code is depicted schematically in the following image.
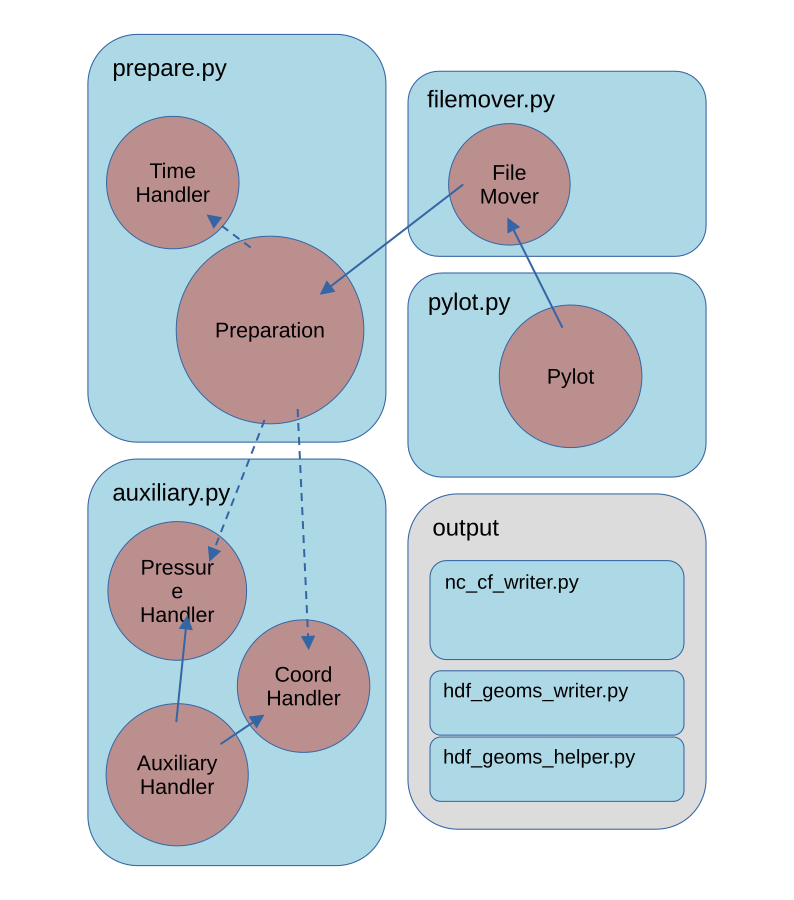
-
The
Pylotclass contains the most top-level functionality for steering PROFAST. The user creates an instance of thePylotclass. This class is inheriting functionality fromFileMoverandPreparationclasses. -
The
Filemoverclass contains the functionality to organize all output files in a file structure. -
The
Preparationclass contains most functionality determining the relevant parameters for the processing. It is the most extensive class. -
The functionality from the
PressureHandler,CoordHandlerandTimeHandlerclasses are called from the Preparation class.
These are the most important functions for steering PROFFAST. The most important functions and interactions are described in the following. Additional code for different output formats located in /output.
prepare.py
Generally the user provides an input file or dictionary with the corresponding parameters for the processing. From this many additional options are derived. This happens in the Preparation class. The following functionality is handled by the Preparation class.
Preparation
- Initialize logging
- Read the input parameters and derive parameters for further processing
- Read corresponding instrument parameters from list
- Search available interferograms/ spectra on disk
- Create a list of dates to be processed
- Generate the input parameters needed for the individual PROFFAST input files
- Handling of coordinates (can be either fixed or variable; for variable coordinates see
CoordHandler) - Interpolate mapfile to local noontime
TimeHandler
- Time zone of the processing and list of dates
- Return
local_time,utc_timeandmeas_timefor a given spectrum or interferogram - Determine local noon of a given day for given coordinates (here, always the local timezone of the location is used, never the daylight saving time)
- Cross-check time consistency
filemover.py
PROFFASTpylot disentangles the output, input and program files. Therefore, files that are written to a location inside the PROFFAST folder by the FORTRAN PROFFAST routines are moved at the end of a run. This sorting of the files is handled by the Filemover class.
Filemover
- Create the output file structure
- Move or copy generated files from the prf folder to the result folder
pylot.py
Here the most top-level functions called by the user are present.
Pylot
- Execute the three PROFFAST routines (preprocess, pcxs and invers)
- Call generation of input files for PROFFAST by filling the corresponding parameters into the respective templates
- Combine results
auxiliary.py
PROFFAST needs accurate pressure data. Pressure values can be read from a file with variable formatting and data frequency. This is handled in auxiliary.py in the class PressureHandler. An internal dataframe is created from the file by prepare_pressure_df(). It can be accessed by Pylot.p_df.
For handling observations with a variable position of the instrument, the coord handler interpolates coord data to the observation time steps.
The AuxiliariyHandler class contains all functionality that is shared between both classes.
AuxiliaryHandler
- Read data and construct dataframe
- Parse datetime column from one or multiple columns
- Interpolate data to specific time
PressureHandler
- Create pressure dataframe as a instance of PressureHandler
- Apply pressure Factors
- Filter Outliers
CoordHandler
- Create coordinate dataframe
- For High-Frequency-Data, average over observation period, otherwise interpolate to observation time
Output
There are four types of output:
- Raw output:
The PROFFAST routines are writing several ASCII-formatted tables, which are placed into theraw_output_proffastfolder. - comb_invparms_<...>.csv
Combines all invparms files and renames the most important to identifiable names (seepylot.pylot.combine_results()) - HDF5 (GEOMS)
HDF5 binary files following the GEOMS conventions are produced byoutput/hdf_geoms_helper.pyandoutput/hdf_geoms_writer.py. These files contain the vertical columns, the column-averaged molar fractions, and the column sensitivities, as well as the prior profiles for CO2, CO, CH4 and H2O. - netCDF4 (cf)
This files contains the same information as the HDF GEOMS files but is delivered in netCDF4 format and follows the cf conventions. It is generated byoutput/nc_cf_writer.py.
PROFFASTpylot - Modules
This page lists the internal methods of all modules, for developing purposes.
PROFFASTpylot consists of four main modules:
-
Pylot: High level functionality.
-
Filemover: Create a consistent file structure from the PROFFAST output.
-
Prepare: Derive all processing options from the input.
-
Auxilary: Read and interpolate the pressure and coordinate data.
-
Output: Write additional output formats.
Pylot
- class pylot.Pylot(input_file, logginglevel='info', external_logger=None, loggername=None)
-
Bases:
FileMoverStart all PROFFAST processes.
- clean_files()
-
After execution clean up the files not needed anymore
- combine_results()
-
Combine the generated result files and save as csv.
- run(n_processes=1)
-
Execute all processes of profast.
Run preporcessing, pcxs, invers. The generated data is moved and merged in a result folder.
- run_inv(n_processes=1)
-
Run inverse.
Loops over localdates, generates the input files and runs invers.
- Parameters: n_processes (int) – If n_processes == 1, run_inv_at is called directly. Otherwise it is called via run_parallel.
- Parameters: n_processes (int) – If n_processes == 1, run_inv_at is called directly. Otherwise it is called via run_parallel.
- run_pcxs(n_processes=1)
-
Run pcxs.
- Loops over local dates and executes the following steps:
-
-
check if the abscos bin file exists already.
-
Interpolate the mapfile. If no mapfile is found for a local date, the date is removed from self.local_dates.
-
generate the input file.
-
run pcxs(in parallel).
-
- Parameters: n_processes (int) – If n_processes == 1, run_pcxs_at is called directly. Otherwise it is called via run_parallel.
- run_preprocess(n_processes=1)
-
Main method to run preprocess.
- run_prf_with_inputfile(prf_inputfile, executable, popen_kwargs={})
-
Run PROFFAST with the given inputfile
- _write_ils_file(program_name, output)
-
Write file containing ILS parameters.
Read all ILS parameters from the spectra file header. Check if the ILS parameters are equal for the processing period.
Filemover
- class filemover.FileMover(input_file, logginglevel='info', external_logger=None, loggername=None)
-
Bases:
PreparationCopy, Move and remove temporary proffast Files.
- check_abscosbin_summed_size()
-
Get size of all abscos.bin files. Give warning if too large.
- delete_abscos_files()
-
Delete the abscos.bin files created by pcxs.
- delete_input_files()
-
Delete the input files for preprocess, pcxs and inv
- handle_pT_VMR_files()
-
Copy or move the pT and VMR files created by pcxs.
If delete-abscosbin_files is True, the pT and VMR are MOVED to the result folder.
If delete-abscosbin_files is False, the pT and VMR are COPIED to the result folder.
They contain the prior information and are therefore an important part or the result. Hence, they are wanted to show up in the result folder in any case.
- init_folders()
-
Create all relevant folders on startup if nonexistant.
Check if relevant proffast folders are existant. Folders to be created: - pT, cal directories - result folder (backup of previous results) - logfiles
- move_input_files()
-
Move the input files for prep., pcxs and inv to result folder
- move_results()
-
Move the gererated files to the result folder.
The invparms_?.dat, job_?.spc and version_?.dat files are searched and moved to the result folder. If files are not found, a warning is printed.
- The colsens.dat are produced by PXCS and
-
-
moved if delete_abscosbin_files is True
-
copied if delete_abscosbin_files is False.
-
This is to ensure that every run has them in the result folder, independent if pcxs was executed or skipped in this run.
Prepare
Auxiliary
class auxiliary.AuxiliaryHandler(description_file,
-
Bases:
objectParent class for reading, interpolating and averaging auxiliary data.
- concat_files(file_list).
-
Read all files from list and concatenate them.
- create_df()
-
Read relevant files and return dataframe.
This function was formerly called _read_subdaily_files().
- create_df_when_date_in_filename()
-
Create dataframe if date information is only given in the file name.
- get_date_list(dates)
-
Return extended, sorted deepcopy of date_list.
interpolate_data_at(df, utc_time, data_key)-
Return interpolated value for given date_key and time stamp.
If time difference greater than threshhold, return None.
- parse_datetime_col(df, date=None)
-
Parse the dataframe for a suitable datetime.
Add the column ‘parsed_datecol’ to the dataframe Depending on the options given, the datetime column is constructed f rom the combination of the separate time and date columns.
- Parameters: df (pandas.DataFrame) – pressure dataframe containing time information in arbitrary format.
- Returns: df (pandas.DataFrame) – with an additional datetime column.
-
- class auxiliary.CoordHandler(coord_type_file, coord_path, dates, logger, static_coords=None)
-
Bases:
AuxiliaryHandlerOrganize variable coordinates from a file for a moving observer.
add_static_coord(df, coord_type)-
Use static coordinates if one of the coordinates is not given.
get_coords_at(utc_time)-
Return coordinates at given utc time.
- interpolate_coords(utc_time)
-
Interpolate coords at a given time.
- Parameters: utc_time – Timestamp
- Returns: oords (list) – Containing interpolated latitude, longitude and altitude. None, if one of the interpolatetd coordinates is None
prepare_coord_df()-
Create dataframe and optinoally add static column.
- class auxiliary.PressureHandler(pressure_type_file, pressure_path, dates, logger, measurement_time=0)
-
Bases:
AuxiliaryHandlerRead, interpolate and return pressure data from various formats.
- get_pressure_at(pressure_time)
-
Return the interpolated pressure at a given time.
If the value is rejected or an interpolation error occured p=0 is returned.The corresponding spectra will not be processed. This is determined in prepare.get_spectra_pT_input().
If the pressure measurements for a whole day is missing, the whole day is deleted from the processing list in pylot.run_inv().
- Parameters: (datetime (pressure_time) – datetime): time in timezone of the pressure file
prepare_pressure_df()-
Read the pressure of a day, from files with a various frequencies.
The dataframe self.p_df is created as a object of the pressure_handler instance Containing the pressure and a datetime column.
The pressure column is multiplied by the pressure_factor given in the pressure input file.
-
Output
-
NetCDF (following cf-conventions)
- class output.nc_cf_writer.NcWriter(path_results)
-
Bases:
objectCreate netcdf files from PROFFAST output following the cf conventions.
- add_avk(ds)
-
Add averaging kernel variables to dataset.
-
Add column with local noot of given date.
This column links the values dependet on time to the values dependent on time_prior.
- add_prior(ds)
-
Add prior variables to dataset.
- create_dataset()
-
Combine all proffast output in one ds.
1. combine all data
2. replace secondary time columns
3. scale to SI units
4. write all metadata and set fill_values
Returns: dataset (xr.Dataset) – dataset following the cf conventions
get_avk_dims()-
Return dict with dims for averaging kernels.
- get_colsens_for(path, species)
-
Return the column sensitivities for a given species.
- get_coords()
-
Read coords from first line of comb_invparms. This is used for the time offsets, recording in multiple time zones is not supported.
- get_files_colsens()
-
Return filelist with all colsens files.
- get_files_prior()
-
Return filelist with all prior (VMR_fast_out.dat) files.
- get_prior(path)
-
Return df with prior values maped on the internal altitude grid.
The values are converto to SI units.
The VMR file has no header. It contains: 0: “Index”, 1: “Altitude”, 2: “H2O”, 3: “HDO”, 4: “CO2”, 5: “CH4”, 6: “N2O”, 7: “CO”, 8: “O2”, 9: “HF”
- get_prior_dims()
-
Return dict with dims for prior variables.
- get_prior_time(cf=True)
-
Return cf times.
- get_time_str()
-
Return string with one year or year range.
- modify_spectrum(ds)
-
Convert spectrum column to cf compatible string type.
- modify_time(ds)
-
Rename, and make cf conform main time column.
- scale_columns(ds)
-
Apply scaling factors to match SI units.
- write_nc(path=None)
-
Write dataset to cf-conform netcdf file.
- class output.nc_cf_writer.PhysicsConstants
-
Bases:
objectClass to store Physics constants.
-
HDF5 (following GEOMS conventions)
- class output.hdf_geoms_helper.GeomsGenHelper(geomsgen_inputfile, geoms_out_path=None)
-
Bases:
object - apply_quality_checks(df)
- defaults = {'DATA_DESCRIPTION': None, 'DATETIME_NOTES': '', 'geoms_out_path': None, 'ils_file': None, 'ils_filelist_warning': False, 'ils_not_in_file_warning': False, 'input_file_name': 'proffastpylot_parameters.yml', 'n_removed_values': 0}
- get_comb_invparms_df(day)
-
Get the complete path to the combined invparms file.
- get_datetimes()
-
Return list of datetimes (local time).
- get_end_date()
- get_start_date()
- mandatory_options = ['prf_res_path', 'geoms_start_date', 'geoms_end_date']
- class output.hdf_geoms_writer.GeomsGenWriter(geomsgen_inputfile, geoms_out_path=None)
-
Bases:
GeomsGenHelper - create_invparms_content(day)
-
Write or overwrite self.df
- generate_GEOMS_at(day)
-
Create a HDF5geoms file for a specific day
- generate_geoms_files()
- get_col_unc(df)
- get_colsens_int(df, day)
- get_colsens_sza(day)
- get_data_content(day)
- get_ils_parameters(local_date)
-
Return ILS Parameters if possible.
ILS parameters are obtained in the following way: 1. if present the values are taken from the generated prf output (since v1.4) 2. else, the values are taken from the preprocess input file 3. else, the values can be taken from a given ils-list (e.g. from the default list shiped with proffastpylot) if the list is given in the geoms input file.
If the values can be obtained from 1. or 2. and additionally from 3. a cross-check is preformed.
- Parameters: day – day to be processed
- Returns: tuple (ME1, PE1, ME2, PE2)
- get_pt_content(day)
- get_start_stop_date_time(day, df)
- get_start_stop_date_time_csv(day)
-
Get start and stop time for each measurement day
- get_vmr_content(day)
- write_air_density(df, ptf, cont)
- write_air_density_src(df, cont)
- write_air_partial(df, ptf, cont)
- write_altitude(df, ptf, cont)
- write_apr(df, ptf, vmr, cont)
-
Write prior total vertical column for each trace gas to the HDF5 file. “XXX_APR”: “XXX.MIXING.RATIO.VOLUME.DRY_APRIORI
- write_apr_src(df, cont)
- write_avk(df, ptf, sen, cont)
- write_col(df, cont)
- write_col_unc(df, cont)
- write_datetime(df, cont)
- write_instr_altitude(df, cont)
- write_instr_latitude(df, cont)
- write_instr_longitude(df, cont)
- write_metadata(day, df)
-
Write metadata to the GEOMS file!
- write_pressure(df, ptf, cont)
- write_pressure_src(df, cont)
- write_solar_angle_azimuth(df, cont)
- write_solar_angle_zenith(df, cont)
- write_source(cont)
- write_surface_pressure(df, cont)
- write_surface_pressure_src(df, cont)
- write_temperature(df, ptf, cont)
- write_temperature_src(df, cont)

